A deep-dive resource covering discovery frameworks, platform architectures, IP valuation, clinical case studies, ROI modeling, regulatory strategy, and the next technology horizon.
Section 1: Why Drug Repurposing Demands a New Analytical Framework
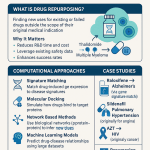
The standard pharmaceutical R&D model has a structural problem that cost-cutting alone cannot fix. A new molecular entity (NME) takes 10 to 15 years to reach the market and costs, on average, more than $2 billion when accounting for the cost of failures. Of every 10,000 compounds that enter preclinical screening, fewer than ten reach human trials. Of those that do, roughly 90% fail, primarily on efficacy or unforeseen toxicity grounds. That math does not improve with incremental process optimization.
Drug repurposing, also called drug repositioning, drug rescuing, or drug redirecting depending on the specific strategic context, offers a structurally different R&D equation. By starting with a compound that already has at least some clinical safety data, a repurposing program skips the most expensive and failure-prone segment of the development timeline. Average development timelines drop to 3 to 12 years. Average costs fall toward $300 million. The probability of clinical success, at roughly 30% for repurposed compounds overall, is approximately three times higher than for NMEs entering Phase I.
The pharmaceutical archives hold thousands of shelved, abandoned, or off-patent compounds with established safety profiles and no approved indication beyond their original target. Sildenafil, developed as a cardiovascular agent and repurposed as Viagra for erectile dysfunction after unexpected trial findings, is the canonical example. Thalidomide, withdrawn in the 1960s after causing severe birth defects as a sedative for morning sickness, was subsequently approved for erythema nodosum leprosum and, later, multiple myeloma. These are celebrated accidents, not replicable strategy.
The shift that matters now is the move from opportunistic discovery to systematic data-driven identification. That shift requires computational tools capable of processing datasets far too large and complex for manual analysis: millions of electronic health records, the entire corpus of published biomedical literature, genomic databases spanning hundreds of thousands of patients, and patent filings across dozens of jurisdictions. Artificial intelligence and machine learning are the only practical tools for this scale of work.
This document covers the full technical and strategic landscape for practitioners who need more than a conceptual introduction. It addresses specific algorithmic architectures, their commercial implications, IP valuation frameworks for the key platform companies, real clinical data from the field’s most important trials, and a structured investment strategy for analysts tracking this space.
Key Takeaways: Section 1
The core economic case for drug repurposing rests on three measurable advantages: a 3x improvement in clinical success rates over NMEs, a cost structure roughly 85% below de novo development, and timeline compression from 12+ years to 3 to 12 years. AI does not change the fundamental logic; it scales the candidate identification process from hundreds of hypotheses to millions, and shifts discovery from human intuition to data-driven pattern recognition. The commercial value unlocked depends entirely on the IP architecture surrounding the repurposed compound, a point that the BenevolentAI/baricitinib case makes starkly clear.
Section 2: The Foundational Logic: Three Discovery Architectures and Their Commercial Ceilings
Before evaluating AI tools, it is worth being precise about what problem they are solving. Drug repurposing research historically follows three distinct strategic logics, each with different starting conditions, different data requirements, and importantly, different commercial ceilings. AI accelerates all three but does not resolve the underlying IP constraints that govern each.
2.1 The Drug-Centric Architecture
The drug-centric approach starts with a specific molecule and asks what else it might treat. The practical manifestation takes several forms. First, systematic analysis of off-label prescribing patterns: physicians routinely prescribe approved drugs outside their labeled indication based on biological rationale or clinical observation, and this real-world usage generates signal that can be formalized through retrospective study. Second, rescue programs for shelved compounds: the pharmaceutical industry has hundreds of compounds that demonstrated acceptable safety profiles in Phase I but were abandoned due to inadequate efficacy in the target indication, and these assets constitute a low-risk starting point because Phase I data already exists. Third, lifecycle management for compounds nearing patent expiration: when a drug’s composition-of-matter patent expires, the originator company faces generic competition on the original indication. A new patented indication, secured before expiration, can extend commercially relevant exclusivity.
The commercial ceiling of the drug-centric approach depends almost entirely on whether the new indication can be protected by a method-of-use patent and whether that patent is strong enough to prevent substitution by the existing generic. For a compound still under composition-of-matter protection, a new indication represents a straightforward expansion. For an off-patent compound, the analysis is more complex and the IP tools available, primarily method-of-use patents and formulation patents, require careful prosecution to be commercially defensible.
2.2 The Disease-Centric Architecture
The disease-centric approach starts with an underserved indication and asks which existing molecules might address it. This logic is particularly relevant for rare and neglected diseases where the market economics of de novo development are commercially unviable. The Orphan Drug Act creates a separate commercial logic for these programs: orphan designation in the U.S. provides seven years of market exclusivity independent of patent status, a significant protection for compounds that might otherwise have weak IP.
The disease-centric approach requires detailed knowledge of the target disease’s biological underpinnings and an ability to scan the existing pharmacopeia for mechanistic overlap. An AI system that has mapped the molecular pathways of Castleman disease, for example, can in principle identify all approved drugs that modulate proteins implicated in those pathways, generating a prioritized candidate list in hours rather than years.
The commercial ceiling here is often limited by patient population size, payer willingness, and the strength of the orphan exclusivity relative to any composition-of-matter protection on the identified compound. A repurposed generic drug with orphan designation but no formulation patent is commercially defensible for seven years in the U.S. but remains exposed in markets with shorter or absent orphan exclusivity periods.
2.3 The Target-Centric Architecture
The target-centric approach is the most scientifically precise. It starts with a validated molecular target: a protein, enzyme, or receptor known to be pathologically relevant in a specific disease. The question is whether an existing drug, approved for a different indication, already modulates that target with sufficient potency and selectivity. This architecture can bridge therapeutically distant indications: a cardiovascular drug that acts on a receptor also implicated in neurological disease becomes a legitimate repurposing candidate once the target connection is established.
The commercial ceiling of the target-centric approach depends on how well the connection between the new target and the known drug can be established as novel and non-obvious. If the target connection was previously described in the literature, a method-of-use patent claim faces a significant prior art challenge. If the AI system is the first to identify the connection, and this can be documented with a clear timestamp, the non-obviousness argument strengthens considerably.
2.4 How AI Changes the Strategic Picture
These three architectures have historically operated in isolation, pursued by separate teams with separate tools. An AI system built on a comprehensive knowledge graph collapses this separation. The same system can simultaneously execute drug-centric queries (what diseases are topologically adjacent to Drug X in the biological network?), disease-centric queries (which existing drugs interact with proteins implicated in Disease Y?), and target-centric queries (what approved molecules bind Target Z with confirmed activity above a defined threshold?). The queries run concurrently, and the results can be cross-referenced to identify candidates that score well across all three frameworks simultaneously.
The other major change is the shift from hypothesis-driven to hypothesis-free discovery. Traditional repurposing requires a biological rationale before the search begins. An AI system trained on electronic health record data from millions of patients can identify a statistically significant association between Drug A and a lower incidence of Disease B without any prior mechanistic hypothesis. The correlation is found first; the biological explanation is sought afterward. This opens a class of repurposing candidates that human researchers would not naturally generate, because the connection is not mechanistically intuitive from current biological knowledge.
Key Takeaways: Section 2
The three repurposing architectures, drug-centric, disease-centric, and target-centric, each have different commercial ceilings determined primarily by IP architecture rather than scientific merit. AI systems built on comprehensive knowledge graphs run all three architectures simultaneously, generating candidates that would not emerge from any single framework in isolation. The hypothesis-free discovery enabled by large-scale EHR analysis is the most scientifically novel contribution AI makes to the field, and it is also the most difficult for competitors to replicate because it requires both the computational infrastructure and access to the underlying patient data.
Section 3: The AI Engine Room: Knowledge Graphs, Deep Learning, Graph Neural Networks, and NLP
Drug repurposing at scale requires four distinct technical capabilities: structured representation of biological knowledge, prediction of novel relationships within that knowledge, reading of unstructured scientific text to keep the knowledge current, and tools that can explain their predictions in terms scientists can act on. Each of these capabilities corresponds to a specific family of AI methods.
3.1 Knowledge Graphs: The Biological Map
A knowledge graph (KG) is a network structure where entities, drugs, diseases, genes, proteins, pathways, symptoms, clinical outcomes, are represented as nodes, and the relationships between them, “inhibits,” “activates,” “is associated with,” “is contraindicated with,” are represented as directional edges. The practical result is a queryable map of biology.
The key metric for a pharmaceutical KG is not size alone but coverage and curation quality. A KG that integrates DrugBank (detailed drug-target binding data), KEGG (pathway and disease ontology data), ChEMBL (bioassay activity data for small molecules), OMIM (genetic disease associations), and the full text of published literature through NLP extraction is qualitatively different from one that combines only public databases. BenevolentAI’s proprietary KG, the system that identified baricitinib as a COVID-19 candidate, contains tens of millions of relationships extracted from scientific publications and curated databases. The density and accuracy of those relationships is the company’s core scientific asset, and the reason the hypothesis about NAK inhibition emerged from their system rather than from manual literature review.
KGs enable a class of prediction called link prediction: given the existing structure of the network, which edges are likely to exist but have not yet been experimentally confirmed? A predicted edge between a drug node and a disease node is, in operational terms, a repurposing hypothesis. The precision of this prediction depends on the quality of the surrounding network structure.
3.2 Deep Learning: Pattern Recognition at Molecular Scale
Deep learning uses multi-layer neural networks to learn features from data without requiring human engineers to specify what those features should be. In drug repurposing, deep learning is applied across a range of tasks, including prediction of drug-target binding affinity, prediction of off-target toxicity, and classification of biological sequence data.
Several neural network architectures are particularly relevant. Recurrent neural networks (RNNs) and convolutional neural networks (CNNs) operate on sequence data, reading SMILES strings (simplified molecular input line entry system, a text-based molecular representation) or amino acid sequences of proteins, and learning the patterns associated with binding activity. These sequence-based models have the practical advantage of not requiring three-dimensional structural data, which is unavailable for many protein targets.
Transformer architectures, originally developed for machine translation and now the basis of large language models, apply a self-attention mechanism that weights the importance of different positions in a sequence relative to each other. When applied to protein sequences, transformer models can predict the functional consequences of amino acid substitutions with accuracy that was unattainable five years ago. AlphaFold2, DeepMind’s protein structure prediction system, used a transformer-based architecture to solve a problem that had resisted the structural biology field for 50 years, predicting the three-dimensional structure of virtually any protein from its amino acid sequence alone.
3.3 Graph Neural Networks: Learning the Shape of Relationships
Graph neural networks (GNNs) extend deep learning explicitly to graph-structured data. Rather than treating a molecule as a flat string, a GNN treats it as a graph where atoms are nodes and chemical bonds are edges. The network learns representations of atoms in the context of their local chemical environment, enabling predictions about molecular properties that reflect actual chemical structure.
When applied to a biomedical knowledge graph, GNNs perform link prediction by learning the structural patterns associated with known drug-disease relationships and extrapolating those patterns to predict unknown ones. The technical advantage of GNNs over simpler embedding methods is their ability to capture higher-order structural information, the way a node is connected to its neighbors, and how those neighbors are connected to their own neighbors, rather than only the direct neighborhood.
GNNs are the primary computational engine behind the repurposing predictions made by platforms like BenevolentAI’s knowledge graph system and Atomwise’s AtomNet. Their performance scales with the density and quality of the underlying graph, which is why data curation strategy directly determines the quality of predictions.
3.4 Natural Language Processing: Turning Literature into Structured Knowledge
Approximately 80% of biomedical data exists as unstructured text: journal articles, clinical trial reports, patent documents, conference abstracts, and electronic health records. NLP systems extract structured information from this text, performing two primary tasks. Named entity recognition (NER) identifies and tags entities in text, drug names, gene symbols, disease terms, protein names, and maps them to standardized ontologies so that “Metformin,” “metformin hydrochloride,” and “1,1-dimethylbiguanide” are recognized as the same entity. Relation extraction identifies the biological relationship expressed in a sentence: “Drug X inhibits Protein Y” is extracted as a directed edge in the knowledge graph.
Modern NLP systems for biomedical text use transformer-based language models, fine-tuned on domain-specific text. BioBERT, SciBERT, and more recent large language models (LLMs) trained on PubMed and patent corpora can extract relationships from new publications as they appear, keeping a knowledge graph current with the literature without manual curation. The practical consequence is that a platform’s KG can grow by thousands of new relationships per week, continuously sharpening the precision of its link predictions.
Patent documents are a particularly valuable NLP target because they contain technical information about mechanism of action, formulation, and synthesis routes that does not always appear in peer-reviewed publications, and the filing date of a patent provides a timestamp that is legally significant for establishing priority.
3.5 Platform Architecture: The Integrated Pipeline
The competitive advantage in AI-driven drug repurposing does not belong to any single algorithm. It belongs to the platform architecture that integrates all four capabilities into a coherent pipeline. The operational sequence is: NLP extracts structured knowledge from literature and patents, populating and continuously updating the KG; GNNs run link prediction across the KG to generate ranked repurposing hypotheses; deep learning models score the top candidates on binding affinity, ADME properties, and predicted toxicity; Explainable AI (XAI) tools provide the biological reasoning behind each prediction in terms that medicinal chemists and biologists can evaluate. Human scientists review the output of this pipeline, prioritize candidates based on scientific and commercial criteria, and commission experimental validation.
This pipeline compresses the candidate identification phase of drug repurposing from two to four years of manual work to weeks of computational work. The bottleneck shifts downstream, to experimental validation and clinical development, where human expertise remains irreplaceable.
Key Takeaways: Section 3
The four core AI technologies, knowledge graphs, deep learning, GNNs, and NLP, each address a distinct technical problem in drug repurposing. GNNs are the most powerful prediction engine for drug-disease link prediction when combined with a high-quality KG. NLP is what keeps the KG current. Deep learning models score individual candidates on molecular properties. No single technology provides competitive advantage; the integrated platform architecture does. The quality of the NLP pipeline and the comprehensiveness of the KG are the hardest components to replicate, which is why both are treated as core IP by platform companies.
Section 4: The Data Stack: Multi-Omics, EHRs, and the Strategic Value of Patent Intelligence
AI algorithms are deterministic given their training data. The quality, breadth, and curation of the data stack is therefore the most important determinant of platform performance, and the most durable source of competitive differentiation.
4.1 Public Bioinformatics Databases: The Necessary but Insufficient Foundation
Public databases form the necessary starting point for any repurposing AI platform. DrugBank provides comprehensive drug-target binding data and ADME profiles. KEGG maps biological pathways and disease ontologies. ChEMBL contains bioassay activity data for more than two million compounds across tens of thousands of assays. PubChem provides structural data for over 100 million compounds. OMIM catalogs the genetic basis of inherited diseases. The Therapeutics Data Commons (TDC), developed at Harvard, packages curated, AI-ready datasets from these and other sources into standardized formats specifically designed for machine learning tasks in therapeutics.
The limitation of public databases is that they are available to everyone. A platform trained exclusively on public data cannot generate predictions that competitors cannot also generate with the same inputs. The commercial differentiation comes from what a platform adds on top of the public foundation: proprietary experimental data, curated clinical data, and the algorithmic quality of the models trained on all of it.
4.2 Multi-Omics Integration: The Biological Context Layer
Multi-omics refers to the integration of several layers of biological measurement data for the same biological system, typically covering genomics (DNA sequences and variants), transcriptomics (RNA expression profiles, indicating which genes are active in a given cell type), proteomics (protein abundance and post-translational modification data), and metabolomics (small molecule metabolite profiles reflecting cellular metabolic state).
Each omics layer provides a different window onto the same underlying biology. A drug that modulates a specific protein target may produce predictable transcriptomic changes, and a platform that has learned the mapping between drug exposure and transcriptomic response across thousands of experiments can use transcriptomic data from a disease context to identify drugs that would push the diseased transcriptome toward a healthy state. This approach, sometimes called transcriptomic signature matching, was pioneered by the Connectivity Map (CMap) project at the Broad Institute and has been extended significantly by AI methods that can handle the high-dimensional data more effectively than traditional statistical approaches.
The commercial value of a proprietary multi-omics dataset comes from two sources. First, it enables more precise biological characterization of disease states and drug mechanisms than public data alone allows. Second, if the dataset was generated through systematic internal experiments (as Recursion Pharmaceuticals generates through its robotic cell imaging platform, which produces hundreds of terabytes of imaging data per year), it is not replicable by competitors without equivalent experimental infrastructure.
4.3 Electronic Health Records and Real-World Data: The Hypothesis-Free Discovery Layer
Electronic health records (EHRs) and healthcare claims databases represent the largest and least exploited data asset in pharmaceutical research. A longitudinal EHR dataset covering millions of patients over a decade contains, in principle, the observational signal from thousands of unintentional drug-disease experiments: patients who happened to take a drug for one condition and whose outcomes for a second condition differed systematically from patients who did not take that drug.
AI models trained on EHR data can identify these signals through pharmacovigilance analysis, survival analysis, and causal inference methods. The hypothesis-free character of EHR discovery is genuinely different from the other data modalities: a statistical association between metformin use and reduced incidence of certain cancers was identified through observational data before any mechanistic explanation existed, and it has since generated a substantial body of prospective research. EHR-based discovery generates the initial signal; mechanistic research and prospective trials provide the validation.
The data access problem here is significant. Large, longitudinal, de-identified patient datasets are held by health systems, insurance companies, and government bodies, not by pharmaceutical companies. Platforms that have established data-sharing agreements with major health systems, such as TriNetX or Flatiron Health, have a structural access advantage that is difficult for new entrants to replicate quickly.
4.4 Patent Data as a Predictive Intelligence Layer
Patent databases are routinely used for freedom-to-operate analysis and competitor monitoring. That is the baseline. The more sophisticated application, and the one most directly relevant to AI-driven repurposing strategy, is using structured patent data as an input to the discovery and prioritization pipeline itself.
A patent document contains the most technically precise publicly available description of a drug’s composition, mechanism, formulation, and manufacturing process. An NLP system applied to the full text of patent filings can extract relationships that do not appear in the peer-reviewed literature, because patents are often filed before scientific publication. The filing date also establishes the earliest documented date of a specific technical insight, which is relevant to both prior art analysis and to documenting the novelty of AI-generated discoveries.
Beyond individual patent analysis, the patterns of patent filing activity provide strategic signal. A cluster of new patent filings around a specific biological target suggests that multiple companies have identified that target as commercially relevant, which can either validate a repurposing hypothesis or signal that the opportunity is becoming crowded. A gap in patent coverage for a specific new indication of an approved drug may represent an uncontested opportunity. A series of patent term extension filings around a specific compound signals that the originator is aggressively defending the asset, which is relevant to commercial risk modeling.
DrugPatentWatch provides structured, searchable data on patent expirations, litigation history (including Paragraph IV filings and inter partes review proceedings), patent term extensions, and regulatory exclusivity periods including orphan drug exclusivity, data exclusivity, and pediatric exclusivity. For a repurposing program that has identified a candidate, this data answers the fundamental commercial viability question: how much time does the program have before the compound faces generic competition on the original indication, and what IP tools are available to create new exclusivity on the new indication?
The most advanced integration goes beyond using patent data as a filter after candidate identification. A training dataset that includes historical patent data alongside clinical outcome data can teach an AI model to distinguish the patent signatures of commercially successful repurposing programs from those that failed despite scientific merit, treating IP architecture as a feature of the prediction model rather than a downstream check.
Key Takeaways: Section 4
The data stack for AI-driven repurposing has four layers: public databases (universal access, no competitive differentiation), proprietary experimental data (high investment, durable advantage), EHR and real-world data (access-dependent, generates hypothesis-free discovery), and patent intelligence (structurally undervalued as a predictive input, not just a compliance check). The competitive differentiation between platforms increasingly rests on the quality of layers two through four, because layer one is equally available to all. Platforms that have systematically integrated structured patent data into their discovery pipeline, not just as a final IP risk screen but as a predictive feature in candidate scoring models, hold a strategic advantage that is not reflected in most public comparisons of AI drug discovery performance.
Section 5: Case Studies With IP Analysis
The four case studies below are chosen for a specific reason: each illustrates a different relationship between scientific success and commercial value capture. Scientific success and commercial value are not the same thing in drug repurposing, and the gap between them is almost always an IP architecture problem.
5.1 BenevolentAI and Baricitinib: Scientific Success, Instructive IP Lesson
The discovery. In late January 2020, the London-based AI company BenevolentAI applied its knowledge graph platform to the emerging SARS-CoV-2 outbreak. The platform queried its biomedical KG for existing approved drugs that could both inhibit viral entry into host cells and suppress the JAK/STAT-driven cytokine cascade implicated in severe COVID-19 pneumonitis. The system identified baricitinib, a JAK1/JAK2 inhibitor approved by FDA for rheumatoid arthritis. The mechanistic logic was dual: baricitinib’s JAK1/2 inhibition could address the cytokine storm, and the system predicted that baricitinib could disrupt viral endocytosis through inhibition of Numb-associated kinases (NAK), specifically AAK1 and GAK. This dual-mechanism hypothesis had not been articulated anywhere in the prior literature.
BenevolentAI published its findings in The Lancet Infectious Diseases in February 2020. The NIH ACTT-2 trial and the COV-BARRIER trial subsequently confirmed that baricitinib, both alone and in combination with remdesivir, reduced mortality in hospitalized COVID-19 patients. FDA granted Emergency Use Authorization for baricitinib in COVID-19 in November 2020, later converted to full approval.
The IP architecture. This is where the case becomes instructive. Baricitinib was invented and patented by Incyte Corporation, with commercialization rights licensed to Eli Lilly. The composition-of-matter patents covering baricitinib’s core chemical structure belong to Incyte and Eli Lilly, not to BenevolentAI. BenevolentAI’s discovery was a method-of-use insight: a new therapeutic application for an existing patented compound. Method-of-use patents for new indications are generally narrower than composition-of-matter patents. They can be circumvented by prescribing the branded drug off-label without infringing the new indication claim, and they face a higher obviousness standard when the compound and the therapeutic area are both known.
BenevolentAI’s public market performance since 2020 reflects this structural constraint. The company’s most famous scientific success generated commercial value primarily for Eli Lilly, not for the company that made the discovery. BenevolentAI correctly identified the utility, but the IP architecture of baricitinib meant that value creation from the new indication accrued predominantly to the patent holder.
The baricitinib case illustrates a general principle in AI-driven repurposing: the quality of the scientific insight does not determine the commercial outcome. The IP architecture does. A platform company that generates repurposing insights for third-party drugs faces a structurally difficult value capture problem unless it secures licensing agreements before publishing, files provisional patent applications on the method-of-use claims before any public disclosure, or moves its business model toward internal development rather than partnering.
IP Valuation Note: Baricitinib (Eli Lilly/Incyte). Baricitinib (Olumiant) generated approximately $720 million in global revenue for Eli Lilly in 2023. The core composition-of-matter patents are held by Incyte under U.S. Patent No. 8,158,616 and related families, with Lilly holding commercialization rights. The drug’s U.S. patent expiry, before potential term extensions, is estimated at 2028, at which point the COVID-19 indication adds incremental method-of-use protection that may delay generic entry in certain markets but is unlikely to sustain branded pricing through full generic substitution. The alopecia areata approval (2022) and ongoing trials in systemic lupus erythematosus represent genuine lifecycle extension opportunities, each creating new method-of-use claims around a compound whose composition-of-matter clock is running.
Key Takeaways: Section 5.1
BenevolentAI’s baricitinib prediction is the field’s most important proof-of-concept. It demonstrated that a computational platform could generate a novel, clinically validated, dual-mechanism repurposing hypothesis for a global health emergency in weeks. The scientific validation is complete. The commercial lesson is equally clear: AI-driven repurposing companies that work on third-party compounds must resolve the method-of-use patent problem before the discovery is public, or they will generate scientific credit without proportionate commercial return.
5.2 Insilico Medicine and Rentosertib (ISM001-055): The First AI-to-Clinic Proof of Concept
The discovery. Insilico Medicine’s platform, Pharma.AI, spans three integrated components: PandaOmics for target identification using multi-omics and literature data; Chemistry42 for generative molecular design; and inClinico for clinical trial outcome prediction. The company pointed this integrated system at idiopathic pulmonary fibrosis (IPF), a progressive fibrotic lung disease affecting an estimated 3 million patients worldwide, with a median survival of three to five years from diagnosis and no currently approved therapy that reverses disease progression.
PandaOmics identified TNIK (TRAF2 and NCK-interacting kinase) as a novel therapeutic target in IPF. TNIK had not previously been validated as an IPF target in the clinical literature. Chemistry42 then generated a library of novel small molecules optimized for TNIK inhibition, and from this library, ISM001-055 was nominated as the lead candidate. The full process from target identification to preclinical candidate nomination took approximately 18 months and required synthesis and testing of approximately 60 to 200 molecules, compared to the thousands typically required in traditional programs. The target identification and early clinical data were published in Nature Biotechnology in March 2024.
The Phase IIa GENESIS-IPF trial, a randomized, double-blind, placebo-controlled study of 71 patients across 22 sites in China, reported in November 2024 that ISM001-055 (now named rentosertib) met its primary safety endpoint across all dose levels. The 60 mg once-daily arm showed a mean FVC improvement of +98.4 mL from baseline over 12 weeks, against a mean decline of -20.3 mL in the placebo group. The Phase IIa results were published in Nature Medicine in June 2025, representing the first peer-reviewed proof-of-concept clinical validation of an AI-designed drug for an AI-discovered target.
The IP architecture. Insilico Medicine’s IP position in the rentosertib program is structurally stronger than BenevolentAI’s position in baricitinib, for a specific reason. TNIK as an IPF therapeutic target was identified by Insilico’s AI system; the target was not previously validated. Rentosertib is a novel molecule generated by the company’s generative chemistry system; it is not a repurposed approved drug. This means Insilico holds both the target validation intellectual property and the composition-of-matter patent on the novel compound, the two strongest categories of pharmaceutical IP.
The company filed patents covering the generative biology approaches in 2018 (granted in 2022), and has continued to file on specific compounds and methods as programs advance. The rentosertib program sits within a broader portfolio of over 30 pipeline assets, 10 of which have received IND approval. The strategic intent is to build a portfolio where the AI platform’s output, novel targets and novel molecules, generates IP that is significantly stronger than what is available in traditional repurposing programs.
Insilico also licenses its Chemistry42 and PandaOmics platforms commercially, creating a software revenue stream that is separate from and potentially more durable than the clinical program revenue. The $1.2 billion partnership with Sanofi, covering discovery of up to six new targets, provides both validation of the platform’s scientific capability and non-dilutive capital that funds the internal pipeline.
IP Valuation Note: Insilico Medicine and Rentosertib. Rentosertib is a first-in-class TNIK inhibitor with a composition-of-matter patent position that Insilico controls directly. If rentosertib reaches approval for IPF, the relevant market is approximately 130,000 to 200,000 patients in the U.S. alone, in a disease where the two currently approved drugs, pirfenidone and nintedanib, generate combined annual revenues exceeding $2 billion. A first-in-class disease-modifying mechanism (TNIK inhibition) that achieves FVC improvement rather than merely slowing decline would be positioned at the premium end of that market. The IP runway from composition-of-matter protection, absent a successful Paragraph IV challenge, likely extends to the mid-2040s given the filing timeline.
Key Takeaways: Section 5.2
Rentosertib represents the strongest commercial IP architecture achievable in AI-driven drug discovery: a novel target identified by AI, a novel molecule generated by AI, composition-of-matter protection owned by the discovering company, and Phase IIa clinical validation. The contrast with the baricitinib case is the entire argument for why platform companies in AI drug discovery are moving toward internal pipeline development rather than partnering on third-party compounds. Scientific capability is a commodity that depreciates. IP is the durable asset.
5.3 Atomwise and DNDi: The Neglected Disease Humanitarian Model
Atomwise’s core technology is AtomNet, a deep learning system for structure-based drug discovery. AtomNet treats each compound as an atomic graph and uses a GNN architecture to predict binding affinity between a small molecule and a protein target with three-dimensional structural data. The platform can screen more than 10 million compounds per day against a given target, compared to 10,000 to 100,000 for high-throughput physical screening.
The collaboration with the Drugs for Neglected Diseases initiative (DNDi) targeted Trypanosoma cruzi, the parasite responsible for Chagas disease, which affects an estimated 6 to 7 million people in Latin America with limited effective treatment options and essentially no commercial R&D investment from major pharmaceutical companies. DNDi identified protein targets in T. cruzi that were considered structurally challenging for traditional screening approaches. Atomwise ran virtual screens of millions of compounds against these targets through its AIMS (Artificial Intelligence Molecular Screen) awards program, a philanthropy mechanism through which the company provides AI-based screening at no cost to academic and nonprofit researchers working on neglected diseases.
The program generated novel hit compounds with confirmed biological activity against T. cruzi, which DNDi is advancing through lead optimization. The commercial value to Atomwise from this collaboration is indirect: scientific publication, platform validation, and reputational positioning that supports commercial partnerships with pharmaceutical companies. The AIMS program has completed over 80 screens across dozens of disease areas.
IP Valuation Note: Atomwise. Atomwise has not yet advanced a proprietary drug candidate to IND stage through its internal pipeline, though it nominated its first AI-driven development candidate, an orally bioavailable TYK2 inhibitor for autoimmune diseases, in 2024. The company’s primary commercial asset is the AtomNet platform itself, which generates revenue through research collaborations with major pharmaceutical companies including Merck & Co., Eli Lilly, AbbVie, and academic institutions. The Hansoh Pharma collaboration, announced at a potential value of $1.5 billion across 11 targets, illustrates the scale of deal value that a mature AI screening platform can attract even before internal programs generate clinical data. The TYK2 inhibitor program, if it advances to Phase II with competitive clinical data, would add composition-of-matter IP to the portfolio and raise the company’s valuation anchor substantially.
Key Takeaways: Section 5.3
The Atomwise/DNDi collaboration demonstrates that AI-driven screening platforms can operate profitably in markets where traditional pharmaceutical investment is absent, using philanthropic mechanisms to generate scientific validation at low marginal cost. The commercial ceiling for a platform company that lacks internal pipeline assets is substantially lower than for one that has converted its platform capability into owned composition-of-matter IP. The TYK2 program represents Atomwise’s attempt to make that transition.
5.4 Every Cure: The Systematic Population Screen Model
Every Cure, co-founded by Dr. David Fajgenbaum, a physician-scientist who treated his own idiopathic multicentric Castleman disease with sirolimus after systematic analysis of his biological data and the existing literature, has built a systematic AI platform for what the organization calls “population-scale repurposing hypothesis generation.” The technical architecture integrates biomedical knowledge graph construction (using NLP over the scientific literature and clinical databases), clinical data analysis (including real-world evidence from patient registries and EHR data), and large language model-assisted hypothesis validation.
The distinguishing feature of Every Cure’s approach relative to commercial platforms is scope: the stated goal is to evaluate every approved drug against every disease in a systematic, ranked fashion, rather than focusing a platform on a specific therapeutic area or target class. Every Cure receives federal funding and operates as a non-profit, which resolves the commercial incentive problem that prevents commercial platforms from investing in programs with small patient populations and limited revenue potential.
The sirolimus story that inspired the organization illustrates the core hypothesis of the program: the existing approved pharmacopeia almost certainly contains effective treatments for diseases that have never been evaluated for those treatments, because systematic evaluation has never been attempted at scale. The barrier is not scientific; it is the absence of commercial incentive. Government and philanthropic funding for a comprehensive AI-driven screen could identify candidates for clinical evaluation across hundreds of rare and neglected diseases simultaneously.
Key Takeaways: Section 5.4
Every Cure represents a fundamentally different organizational model: systematic population-scale hypothesis generation funded by philanthropy and government rather than commercial capital. The IP implications are deliberately different from commercial platforms: discoveries made by Every Cure are intended to be shared widely to facilitate development by whoever has the capability and interest, rather than to be captured by a single entity. For commercial investors, this model is relevant primarily as a source of validated repurposing hypotheses that may be available for clinical development by third parties, particularly in rare disease indications with orphan drug exclusivity potential.
Section 6: The Economics: Timeline Compression, Cost Reduction, and Success Rate Modeling
The economic case for AI-driven repurposing rests on three quantifiable improvements relative to de novo drug development: timeline compression, cost reduction, and improved probability of clinical success. Each is measurable, but each requires precision about the baseline being compared.
6.1 Timeline and Cost Baselines
A de novo NME development program runs 10 to 15 years from target identification to approval, at an average fully-loaded cost, including the cost of failures, exceeding $2 billion. Drug repurposing inherently compresses this timeline to 3 to 12 years, with average costs closer to $300 million, by skipping extensive Phase I safety work and leveraging existing manufacturing and formulation knowledge. AI adds a further compression of the candidate identification phase, which in traditional programs consumes two to four years of iterative hypothesis generation and experimental testing. An AI platform can generate and score a candidate list in weeks, with experimental validation taking months rather than years.
The claim that AI can reduce overall development timelines by 50% is plausible but requires a precise scope. AI reduces the discovery and early development phases dramatically. It has less direct impact on Phase III trial duration, which is governed by enrollment rates and the natural history of the disease, and on the FDA review clock. The appropriate benchmark for AI-specific savings is the preclinical discovery and candidate selection phase, where the compression from two to four years to weeks is well documented. The total program timeline benefit is real but smaller than headline claims suggest when Phase II and III timelines are included.
6.2 The Success Rate Distribution
The 30% overall clinical success rate for repurposed drugs, versus 10 to 12% for NMEs entering Phase I, is accurate but masks a wide distribution. Within-class repurposing, moving a drug from one cancer indication to another, has a reported success rate of approximately 67%. Cross-therapeutic area repurposing, moving a cardiovascular drug to a neurological indication, drops to approximately 2 to 5%. Drug rescue programs, attempting to develop a compound that previously failed in a clinical trial, have a reported success rate of approximately 9%.
AI’s greatest economic value is in improving cross-therapeutic area repurposing, because this is where the success rate is lowest and the improvement potential is greatest. A within-class repurposing program does not need AI to achieve a 67% success rate; conventional medicinal chemistry reasoning is sufficient. A cross-therapeutic area program that uses AI to identify and validate a novel biological connection starts with better prior probability than a program initiated from intuition alone.
Early clinical stage data from AI-native biotech companies shows Phase I success rates of 80 to 90% for their AI-designed or AI-selected candidates, compared to approximately 10% for traditional methods. This figure reflects selection bias, because AI companies are selective in what they advance, and small sample size, because the number of AI-originated drugs that have reached clinical trials is still small. But the direction of the effect is consistent with the hypothesis that AI-improved candidate selection materially raises the probability of clinical success.
6.3 The Market Structure
The overall drug repurposing market is growing at a compound annual growth rate (CAGR) of approximately 5.4%, projected to reach $59 billion by 2034. The AI-in-drug-discovery market, which includes but is not limited to repurposing applications, is growing at 10 to 30% CAGR depending on the market research source, with market size projections reaching $8 billion to $16 billion by the early 2030s.
The divergence in growth rates between the overall repurposing market and the AI-driven segment of drug discovery is the key strategic signal. AI is not growing proportionally with the underlying drug repurposing market; it is growing faster because it is expanding the addressable repurposing opportunity by making previously intractable problems tractable. Over 90% of large pharmaceutical companies report active investment in AI capabilities for drug discovery and development.
McKinsey’s estimates of $60 to $110 billion in potential annual value creation for the pharmaceutical industry from AI are plausible in aggregate but should be understood as an upper bound on a long time horizon, not a near-term achievable figure. The realization of this value depends on clinical trial success rates that remain uncertain, regulatory approval timelines that remain long, and commercial adoption of AI-designed drugs that has so far been measured in dozens of clinical programs rather than hundreds.
Key Takeaways: Section 6
The headline economic claims for AI in drug repurposing are broadly correct in direction but require precision to be analytically useful. AI’s most defensible economic contribution is in compressing the candidate identification phase and improving cross-therapeutic area repurposing success rates. The Phase I success rate data for AI-designed drugs (80 to 90%) is preliminary and subject to selection bias but consistent with the theoretical prediction that better candidate selection translates to fewer clinical failures. The market growth data confirms that pharmaceutical investment in AI is accelerating faster than investment in conventional repurposing methods, and this divergence will compound over time.
Section 7: Investment Strategy: Where Value Accrues in AI-Driven Repurposing
This section is written for institutional investors and corporate development teams evaluating positions in the AI drug repurposing space.
7.1 The Platform Company Valuation Problem
AI drug discovery platform companies are valued on a combination of platform capability metrics and clinical pipeline value. The fundamental analytical challenge is that platform capability is not directly observable; investors must infer it from proxy metrics including number of partnerships, deal value, number of IND-approved programs, and platform licensing revenue. Clinical pipeline value is more conventional to model but remains highly uncertain at early stages.
The baricitinib case illustrates the core risk in platform company valuation: scientific validation does not automatically translate to IP ownership, and IP ownership is the prerequisite for commercial value capture. A platform company that generates 10 validated repurposing hypotheses per year but licenses all of them to third parties on milestone-and-royalty terms has a structurally limited earnings ceiling. A company that retains ownership of novel targets and novel molecules, as Insilico is doing with the rentosertib program, has a fundamentally different earnings profile once those programs reach late-stage trials.
The distinction between “AI-assisted discovery for a third party” and “AI-generated novel IP retained internally” is the most important analytic dimension for valuing platform companies. Recursion Pharmaceuticals, BenevolentAI, Insilico Medicine, Exscientia (now merged with Recursion), and Atomwise each represent different positions along this spectrum, and their valuations should reflect these structural differences.
7.2 Key Value Inflection Points
For internal pipeline programs, the conventional biopharmaceutical value inflection points apply: IND filing, Phase I safety readout, Phase IIa proof-of-concept efficacy signal, Phase III enrollment completion, and NDA submission. The Insilico/rentosertib program has already passed the Phase IIa proof-of-concept gate, which is the highest-value milestone for a pre-approval-stage asset. The pivotal trial design and enrollment timeline now determine the next major inflection.
For platform licensing businesses, the relevant inflection points are different: first partner drug to Phase II using the platform’s candidate identification output, demonstrated improvement in partner Phase I success rates compared to historical baseline, first platform-generated NDA approval (regardless of who holds the approval), and platform licensing fee growth rate.
7.3 The IP as Asset Framework
Every repurposing program should be analyzed through an IP-as-asset framework before investment. The relevant questions are: who owns the composition-of-matter patent on the compound? Who owns the method-of-use patent on the new indication? What is the remaining patent term, accounting for any Patent Term Extension (PTE) under Hatch-Waxman for the time the drug spent in clinical development and regulatory review? Is there orphan drug exclusivity, and if so, in which jurisdictions? Is there data exclusivity (five years for small molecules, twelve years for biologics under U.S. law)?
A repurposing program built on a compound with expired composition-of-matter protection and no patent term extension remaining is commercially viable only if a new method-of-use patent can be obtained and defended, the new indication qualifies for orphan exclusivity, a novel formulation can be developed that commands a premium and is protected by formulation patents, or the program can generate an NDA for a rare pediatric disease and earn a priority review voucher (PRV), which can be sold for $100 million or more.
7.4 Sector Positioning
For a diversified position in AI-driven drug repurposing, a portfolio approach across three layers makes sense. The platform layer includes companies that have built and validated AI discovery infrastructure, with revenue from platform licensing providing some near-term cash flow regardless of clinical outcomes. The pipeline layer includes companies with internal AI-generated programs in Phase I or Phase II, where proof-of-concept clinical data can be independently evaluated. The enabler layer includes data and intelligence providers whose value grows with the overall adoption of AI in drug discovery, including specialized patent intelligence platforms that are becoming structurally embedded in the due diligence process.
Platform intelligence providers like DrugPatentWatch sit in the enabler layer. Their value in AI-driven repurposing is not as a discovery tool but as a commercial viability filter and a predictive signal generator for IP trajectory, which makes them infrastructure for the field rather than a bet on any single program.
Key Takeaways: Section 7
The single most important analytic distinction in evaluating AI drug repurposing investments is between companies that generate AI-assisted insights for third-party compounds versus companies that retain IP ownership over AI-generated novel molecules and targets. The former business model generates scientific value more reliably than commercial value. The latter model creates the conditions for conventional pharmaceutical asset valuation once clinical data is available. Apply the IP-as-asset framework to every repurposing program: composition-of-matter protection, remaining patent term, method-of-use claim strength, orphan exclusivity, and data exclusivity status determine the commercial ceiling more reliably than any clinical hypothesis about disease biology.
Section 8: Barriers: Explainability, Data Bias, Talent, Regulatory Risk, and IP Complexity
A candid analysis of the field requires acknowledging the barriers that remain significant. These are not theoretical concerns; they are operational constraints that have slowed adoption and contributed to program failures.
8.1 The Explainability Requirement
Many high-performing deep learning models, particularly deep GNNs applied to large biomedical knowledge graphs, produce predictions without generating human-interpretable reasoning. A biologist receiving a top-ranked repurposing candidate from a GNN-based system needs to know which features of the biological network drove the prediction before committing laboratory resources to validation. Regulators need a mechanistic explanation of a drug’s proposed mode of action before approving it. Neither requirement is satisfiable by a prediction confidence score alone.
Explainable AI (XAI) tools address this problem by providing, alongside the prediction, a human-interpretable explanation of the reasoning. For knowledge graph-based predictions, this typically takes the form of a semantic subgraph: a visualization of the chain of biological relationships, from the drug through intermediary proteins and pathways, to the predicted disease connection, that most influenced the model’s output. The rd-explainer system, developed for rare disease repurposing applications, generates exactly this type of output.
XAI is not a research-stage concern in 2025; it is an operational requirement. FDA and EMA have both issued guidance and discussion papers emphasizing the importance of model transparency and the need for scientifically valid mechanistic rationale in AI-assisted drug development submissions. Companies that have built XAI capabilities into their platforms from the beginning have a regulatory submission advantage over those that treat explainability as a post-hoc addition.
8.2 Data Quality and Algorithmic Bias
The most pernicious data problem in AI-driven drug repurposing is not missing data but biased data. Clinical trial populations have historically been predominantly male and predominantly of European ancestry. The genomic databases that form the foundation of many repurposing knowledge graphs reflect this demographic skew. A model trained primarily on data from one demographic group will generate predictions that may have systematically lower accuracy for other groups, potentially exacerbating existing health disparities.
Compound this with publication bias (positive trial results are published at much higher rates than negative results, creating an optimistically skewed picture of drug efficacy in the literature that NLP systems extract from) and assay bias (activity data in ChEMBL and PubChem reflects the compound classes and target families that researchers have historically been interested in, not a random sample of all possible biological space), and the result is a training data environment that is structurally biased in multiple directions.
Mitigation requires proactive data auditing, deliberate construction of training datasets from diverse sources, and continuous post-deployment monitoring of model performance across demographic subgroups. This is technically tractable but resource-intensive, and many platforms have not yet implemented it systematically.
8.3 Talent: The Bilingual Scientist Problem
A 2024 McKinsey analysis identified a 46% skills gap as a primary barrier to AI adoption in pharmaceutical R&D. The shortage is specific: scientists who are fluent in both computational methods and biological domain knowledge. A data scientist who can build a GNN but does not understand the biology of the knowledge graph they are training it on will generate predictions that are mathematically sound and biologically irrelevant. A biologist who understands the target biology but cannot evaluate the assumptions embedded in a deep learning model cannot effectively oversee the AI’s output.
The talent problem is compounding because pharmaceutical companies are competing for this expertise against technology companies that can offer higher salaries, more flexible work environments, and the cachet of working on foundational AI technology rather than a specific therapeutic application. Pharmaceutical companies retain the advantage of domain-specific datasets and regulatory knowledge that technology companies cannot easily replicate, but this advantage does not fully offset the compensation differential in recruiting.
The organizational structure that most consistently resolves this problem is the integrated, co-located, cross-functional team where computational scientists and experimental biologists share office space and project ownership from the beginning of a program. Siloed hand-offs, where a data science team generates predictions and throws them over the wall to a biology team, consistently produce slower programs with higher rates of computational-experimental misalignment.
8.4 The Evolving IP Landscape for AI-Generated Discoveries
The question of who can be named as an inventor on a patent for a drug discovered by an AI system is unsettled in most major jurisdictions. The U.S. Patent and Trademark Office has maintained that inventors must be natural persons. The Federal Circuit affirmed this position in Thaler v. Vidal (2022). The European Patent Office has reached the same conclusion in the DABUS cases. As AI systems take on a larger role in the actual generation of novel molecules and the identification of novel targets, the legal question of inventorship will require resolution through either judicial or legislative action.
In the interim, the practical IP strategy for companies whose AI systems identify novel molecules or targets is to ensure that the human scientists who guided the AI’s development, who specified the biological problem, who curated the training data, who validated the predictions, and who designed the experimental confirmation are named as inventors. The AI system is treated as a sophisticated tool analogous to a mass spectrometer, which also generates data that humans then interpret into patentable discoveries.
Paragraph IV filings against method-of-use patents covering repurposed drugs are a specific vulnerability. A method-of-use patent covering a new indication for a compound whose composition-of-matter patent is expired faces a Paragraph IV challenge from generic manufacturers at the earliest possible opportunity. The “carve-out” or “skinny label” strategy, where a generic manufacturer seeks approval for the original indication only and explicitly excludes the patented new indication from its labeling, is a standard generic industry tactic that limits but does not eliminate the commercial exposure from method-of-use patents.
Key Takeaways: Section 8
The four principal barriers to AI-driven drug repurposing are addressable but not trivial. XAI is an operational requirement, not a feature: platforms without it cannot generate regulatory-grade mechanistic rationale. Demographic bias in training data is a systemic risk that requires proactive management rather than passive monitoring. The talent shortage for bilingual computational-biological scientists is a 5 to 10 year structural constraint. The AI inventorship question will be resolved by legislation or judicial action, but the practical interim strategy is robust human involvement in the inventive process and clear documentation of that involvement at each stage.
Section 9: The Next Technology Horizon: Generative AI, Quantum Computing, and N-of-1 Medicine
9.1 Generative AI: From Finding Drugs to Designing Them
The distinction between drug repurposing and de novo drug design is becoming less meaningful as generative AI matures. Traditional repurposing finds an existing molecule and asks what else it might do. Generative AI can take an existing molecule as a starting point and generate a family of structurally related compounds, each computationally optimized for improved binding affinity, better metabolic stability, reduced off-target toxicity, and more favorable ADME properties relative to the parent compound.
Insilico Medicine’s Chemistry42 platform, the system that generated rentosertib, is an operational example of this capability. The Pharma.AI workflow begins with target identification (PandaOmics), uses Chemistry42 to generate novel molecules optimized for that target, then uses inClinico to predict clinical trial success probability. The generative step produces candidate molecules that do not exist in any compound library, giving the company composition-of-matter patent rights over the new molecules rather than method-of-use rights over existing ones.
Generative Adversarial Networks (GANs) and Variational Autoencoders (VAEs) learn the statistical properties of known bioactive molecules and can generate new molecules that satisfy specified property constraints, including molecular weight, LogP (lipophilicity), predicted binding affinity, and predicted synthetic accessibility. Diffusion models, the same architecture behind image generation systems, are now being applied to three-dimensional molecular structure generation and protein-ligand co-design.
The commercial consequence is that generative AI transforms repurposing into a hybrid workflow: identify the biological target and approximate binding pharmacophore using repurposing logic and knowledge graph analysis, then use generative chemistry to design a novel, patentable molecule optimized for that specific application. The result is stronger IP than a pure repurposing strategy and faster development than a pure de novo strategy.
9.2 Quantum Computing: Simulating Molecular Physics at Accurate Scale
Classical computing approximates the quantum mechanical interactions that govern drug-protein binding. These approximations are necessary because exact quantum mechanical calculations for systems larger than a few dozen atoms are computationally intractable on conventional hardware. The approximations introduce errors in predicted binding energies that compound across the development pipeline: a compound selected based on an inaccurate predicted binding affinity may fail expensive wet-lab and in vivo experiments that could have been avoided with more accurate predictions.
Quantum computers operate on qubits that can exist in superposition and become entangled, enabling calculation strategies that are fundamentally impossible for classical bits. For drug discovery, the most relevant application is quantum chemistry simulation: calculating the electronic structure and intermolecular forces in a drug-protein binding complex with accuracy that classical methods cannot achieve. Companies including Quantinuum, IBM Quantum, IonQ, and Google Quantum AI are actively working on quantum chemistry algorithms applicable to molecular systems relevant to drug discovery.
Current quantum hardware is not yet fault-tolerant at the scale needed for practically useful drug discovery calculations; this is the field’s honest assessment in early 2025. The near-term value is in hybrid quantum-classical algorithms that use quantum hardware for specific calculation steps where it outperforms classical alternatives. The 5 to 10 year trajectory is toward fault-tolerant quantum computers capable of accurate binding energy calculations for drug-sized molecules, at which point the improvement in predictive accuracy would reduce the failure rate of compounds advanced to expensive experimental testing.
9.3 Multi-Modal Integration: The Digital Patient Model
The prediction models deployed in drug repurposing today are primarily trained on molecular and genomic data. The clinical reality of drug response involves tissue architecture, immune context, metabolic state, and dynamic physiological interactions that are not captured in these modalities alone. Multi-modal models that integrate imaging data (MRI, PET, histopathology), sensor data from wearables, spatial transcriptomics (which provides gene expression data in the context of tissue structure rather than averaged across a tissue sample), and conventional genomics provide a richer and more clinically predictive representation of disease biology.
The technical challenge in multi-modal integration is alignment: different data types have different resolutions, different noise characteristics, and different temporal dynamics. A genomic sequence is static; a wearable-sensor dataset changes by the minute. Integrating these requires sophisticated representation learning that can find common structure across incompatible data formats. Transformer-based architectures with cross-modal attention mechanisms are the current state of the art for this problem.
9.4 N-of-1 Personalized Repurposing: The Long-Term Trajectory
Drug repurposing in its current form produces new indications for diseases defined by diagnostic categories. A repurposed drug is approved for “treatment-resistant depression” or “HER2-positive breast cancer,” categories that still aggregate millions of patients with meaningfully different molecular disease drivers.
The long-term trajectory, enabled by multi-modal integration, more precise genomic characterization, and AI models of sufficient complexity, is toward N-of-1 repurposing: identifying the optimal existing compound or combination of compounds for a specific patient’s unique molecular disease profile. A patient with treatment-refractory cancer whose tumor has been characterized by whole-genome sequencing, RNA-seq, proteomics, and imaging could in principle have their individual molecular profile used as input for an AI screen of all approved drugs and combinations, generating a ranked list of candidates predicted to be effective for their specific tumor biology.
This model requires solving problems that are not yet solved: interpretable multi-modal disease representation, validated causal models of drug response at the individual level, and clinical trial infrastructure designed for single-patient studies. The regulatory pathway for N-of-1 medicine is also undeveloped. But the direction of the technology trajectory, increasing data richness at the individual patient level combined with increasingly powerful AI models, points consistently toward this destination.
Key Takeaways: Section 9
Generative AI is collapsing the distinction between repurposing and de novo discovery, enabling the design of novel molecules optimized for repurposing-identified targets and producing composition-of-matter IP rather than method-of-use IP. This is the most commercially consequential technical development in the field in the near term. Quantum computing is a longer-horizon development that will improve the accuracy of molecular binding predictions and reduce late-stage failures, but fault-tolerant quantum hardware at the required scale is 5 to 10 years away. N-of-1 personalized repurposing is the long-term trajectory of the entire field, enabled by the convergence of multi-modal patient data and increasingly powerful AI models of drug response.
Section 10: Consolidated Key Takeaways and Investment Strategy Summary
10.1 The Ten Points That Matter Most
Drug repurposing achieves a roughly 30% overall clinical success rate versus 10 to 12% for NMEs, with a cost structure roughly 85% lower than de novo development. The variation within that 30% figure is large: within-class repurposing achieves 67%, while cross-class repurposing achieves 2 to 5%. AI’s primary economic value is in improving cross-class repurposing, which is the hardest category and has the largest potential gain.
The AI architecture that drives the most powerful repurposing predictions is the integrated pipeline: NLP for knowledge graph construction, GNNs for link prediction, deep learning for candidate scoring, and XAI for prediction explanation. No single algorithm component is competitively decisive. The integrated pipeline quality is.
The data stack determines platform performance more reliably than algorithmic sophistication. Public databases provide a common floor. Proprietary multi-omics experimental data, curated EHR datasets, and systematically integrated patent intelligence each provide differentiating capability that is not available to all competitors.
The baricitinib case is the field’s most important IP lesson: generating a correct scientific insight about a third-party compound creates scientific credit and regulatory impact without IP ownership. Commercial value accrues to the patent holder. Any AI-driven repurposing program must resolve the IP architecture question before scientific findings are published.
Rentosertib/ISM001-055, published in Nature Medicine in June 2025, is the first peer-reviewed proof-of-concept clinical validation of a drug discovered and designed by AI for an AI-identified target. It establishes a composition-of-matter IP model that is structurally superior to method-of-use repurposing for commercial value capture.
Explainable AI is an operational requirement for clinical and regulatory adoption, not an optional feature. Platforms built without XAI capability cannot generate mechanistic rationale sufficient for regulatory submission.
Algorithmic bias in AI repurposing models is a specific scientific risk, not just an ethical concern. A model trained predominantly on data from one demographic group will generate systematically less accurate predictions for other groups. This translates to clinical failures that could have been predicted and avoided.
Generative AI is converting the repurposing opportunity into a composition-of-matter IP opportunity by enabling the design of novel molecules optimized for repurposing-identified targets. This is the most important near-term commercial development in the field.
Patent intelligence, specifically structured data on patent expiration schedules, Paragraph IV litigation history, patent term extensions, and orphan exclusivity periods, is the most underutilized decision input in repurposing candidate prioritization. It should be integrated as a predictive feature in candidate scoring models, not applied as a post-hoc commercial risk filter.
The talent gap for bilingual computational-biological scientists is a 5 to 10 year structural constraint on AI adoption across the pharmaceutical industry. Companies that have resolved this through integrated team structures and cross-training programs have execution advantages that are observable in program velocity.
10.2 Strategic Investment Positioning
For institutional investors, the AI drug repurposing space offers three distinct risk-return profiles. Companies with AI-generated internal pipelines in Phase II or later, where proof-of-concept data is available for independent evaluation, offer conventional biopharmaceutical risk-return with an AI-derived development cost and timeline advantage. Companies with AI platform businesses generating partnership and licensing revenue offer technology sector valuation dynamics with pharmaceutical sector correlation risk. Patent intelligence and data infrastructure providers offer lower risk, lower return positions that are structurally insulated from individual clinical trial outcomes.
The most attractive risk-adjusted positions are companies where the AI platform has been validated by Phase II clinical data for internally-owned programs, the IP portfolio covers novel targets and novel molecules under composition-of-matter protection, and platform licensing revenue provides a cash flow base independent of clinical trial outcomes. This combination exists in nascent form in the current market and will become more common as the field’s first wave of AI-generated drugs completes Phase II and enters Phase III.
Frequently Asked Questions
What is the right starting point for a mid-sized pharmaceutical company building its first AI repurposing capability?
Start with the disease area you know. The highest-probability early win comes from pointing AI tools at a therapeutic area where your organization has deep biological expertise. Use public databases and commercially available AI platforms rather than building from scratch. The key internal investment is not in proprietary algorithms but in data quality: curating a high-quality disease-specific multi-omics dataset and integrating structured patent intelligence for your target indication. A focused disease-centric program with excellent internal domain expertise is more likely to produce a near-term validated candidate than a broad platform build without biological grounding.
How does AI change the role of medicinal chemists and biologists in an R&D organization?
The role shifts from manual hypothesis generation and iterative experimental screening to hypothesis evaluation and experimental validation. A medicinal chemist in an AI-enabled organization spends less time proposing candidate molecules and more time evaluating the chemical tractability, synthetic accessibility, and structural novelty of AI-generated candidates. The biological domain expertise required to evaluate AI predictions, to identify which predicted drug-disease connections are biologically plausible and which are computational artifacts, becomes more valuable, not less. The scientist who can translate between the language of data science and the language of biology is the organizational asset that will determine program velocity.
How should method-of-use patents be evaluated for repurposed drugs competing against generic versions of the original compound?
The commercial viability analysis requires four components: patent claim scope (how precisely does the claim define the new indication, and can it be worked around by labeling the original indication only?), generic carve-out risk (can generic manufacturers obtain approval using a skinny label that excludes the patented indication?), prescribing behavior (will physicians substitute the generic off-label for the patented indication, and can this be blocked practically?), and payer dynamics (will formularies require step therapy through the generic before authorizing the branded product for the new indication?). Method-of-use patents for repurposed drugs are commercially viable primarily when the new indication is clearly differentiated from any existing generic indication, orphan exclusivity is available, or a formulation patent creates a distinct branded product that cannot be substituted by the existing generic form.
What regulatory submission strategy should companies use for AI-generated drug development evidence?
FDA’s current guidance on AI in drug development, most recently articulated in the agency’s 2021 discussion paper and subsequent draft guidance, emphasizes a risk-based framework focused on model validation, transparency, and data integrity. The practical submission strategy should include full documentation of the training data, including its sources, curation methodology, and known limitations; model validation studies demonstrating performance on held-out data and, where possible, prospective predictions that were subsequently confirmed experimentally; and XAI outputs that provide human-interpretable mechanistic rationale for each key prediction. Engage with FDA’s Emerging Technology Program early for AI-novel development approaches; the agency has been receptive to pre-submission meetings on AI methodology.
What is the single most common organizational failure in AI drug discovery programs?
Treating the AI program as a data science project rather than an integrated R&D strategy. Programs that fail to integrate computational scientists with biologists, chemists, and clinicians from day one consistently produce models that are technically functional and biologically misaligned. The data scientists do not understand what biological questions matter most; the biologists do not understand what data is available or what the model can and cannot do. The organizational fix is co-location and shared project ownership from project initiation, with equal leadership weight given to computational leads and biological leads. Programs with this structure consistently outperform siloed approaches in time to IND filing and Phase I success rate.
Key sources include Nature Medicine (June 2025), Nature Biotechnology (March 2024), BenevolentAI publications, Insilico Medicine clinical disclosures, DNDi program reports, Precedence Research market analysis, and structured patent intelligence from DrugPatentWatch.
Copyright notice: This analysis draws on publicly available scientific publications, company disclosures, and market research. All referenced clinical data, patent information, and market figures are from primary or established secondary sources cited within the text.