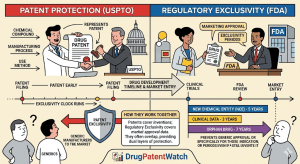
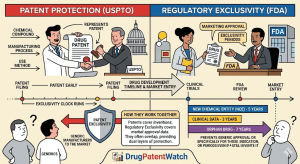
Designing Around the Picket Fence: A Strategic Guide to Freedom-to-Operate for Generic Drugs
An investigative analysis of patent thickets, design-around strategies, and competitive intelligence for pharmaceutical generics manufacturers The Wall Every Generic Manufacturer […]